SARS-Cov-2 Structome – The Viral Machinery behind COVID-19
The structure of all mature proteins from SARS-CoV-2 were predicted using an ensemble workflow that combines the state-of-the-art methods of protein structure prediction. These structures were used as the basis for an extensive exploration of molecular differences between SARS-CoV-1 and SARS-CoV-2 that may explain pathogenicity divergence.
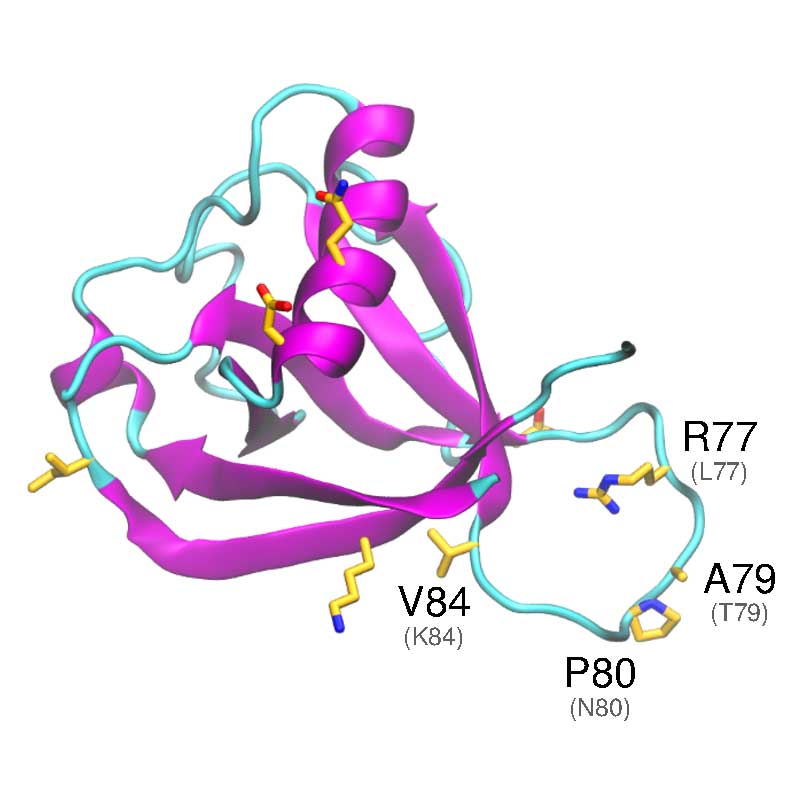
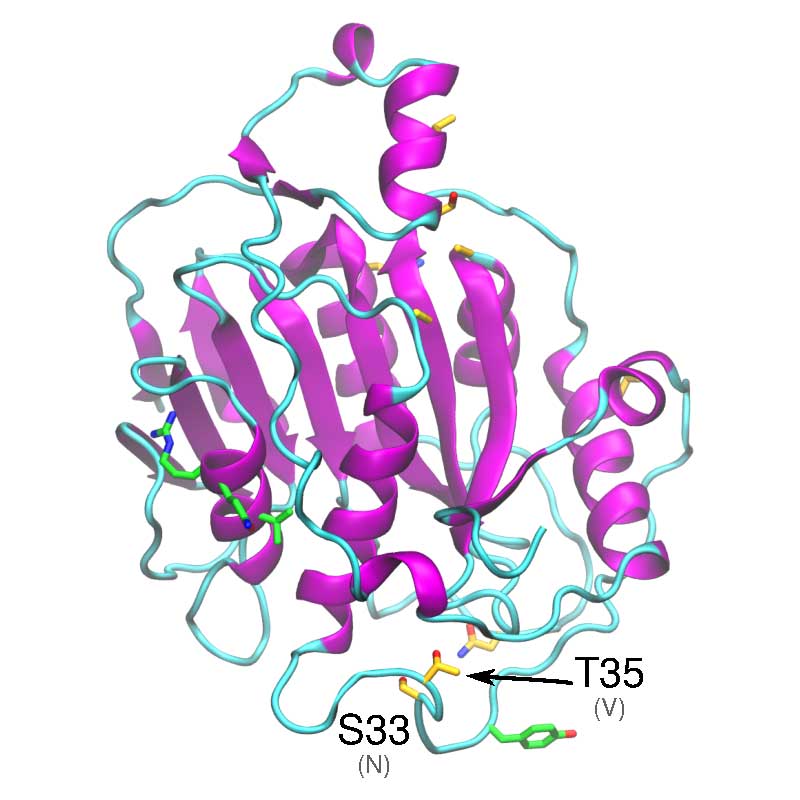
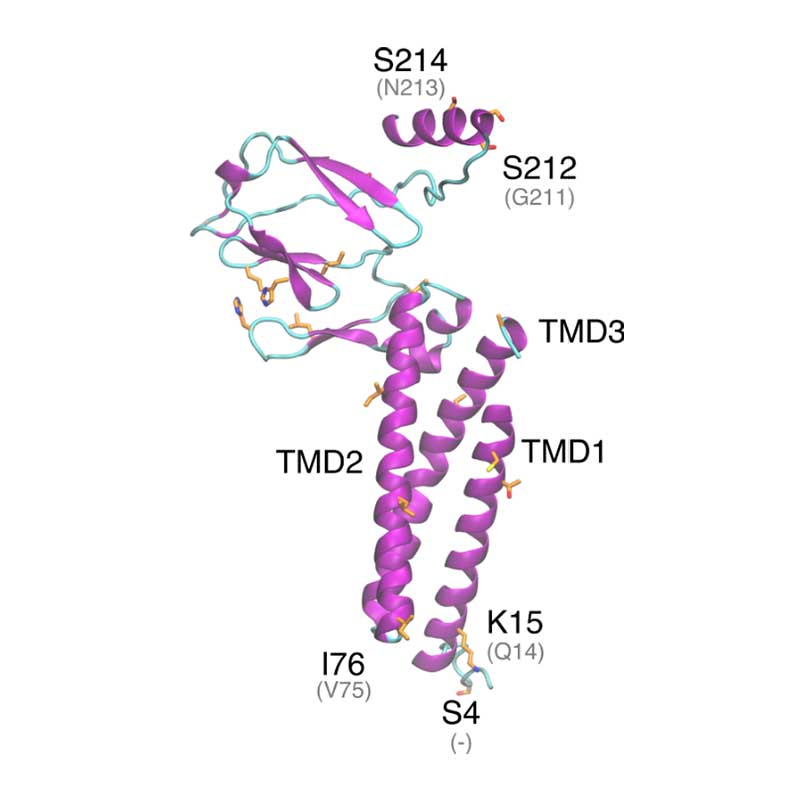
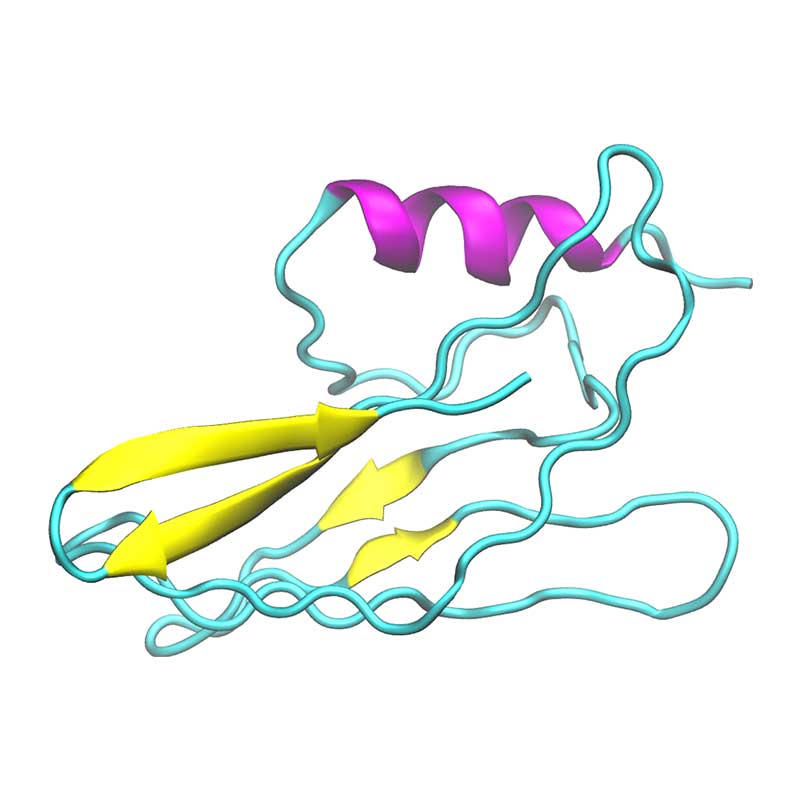
In vitro analyses have found that SARS-COV-1 nsp1 may disrupt the host interferon defense response, by potentially affecting the downstream defense signaling. The nsp1 protein has been shown to bind the 40S Ribosomal subunit, which has been associated with degradation of host mRNA, and suppression of host mRNA translation, leaving the viral RNA unaffected. SARS-CoV- 1 vs SARS-CoV-2: Variations in Loop 3-4 may influence the ability of SARS-CoV-2 nsp1 to disrupt host-translational activity.
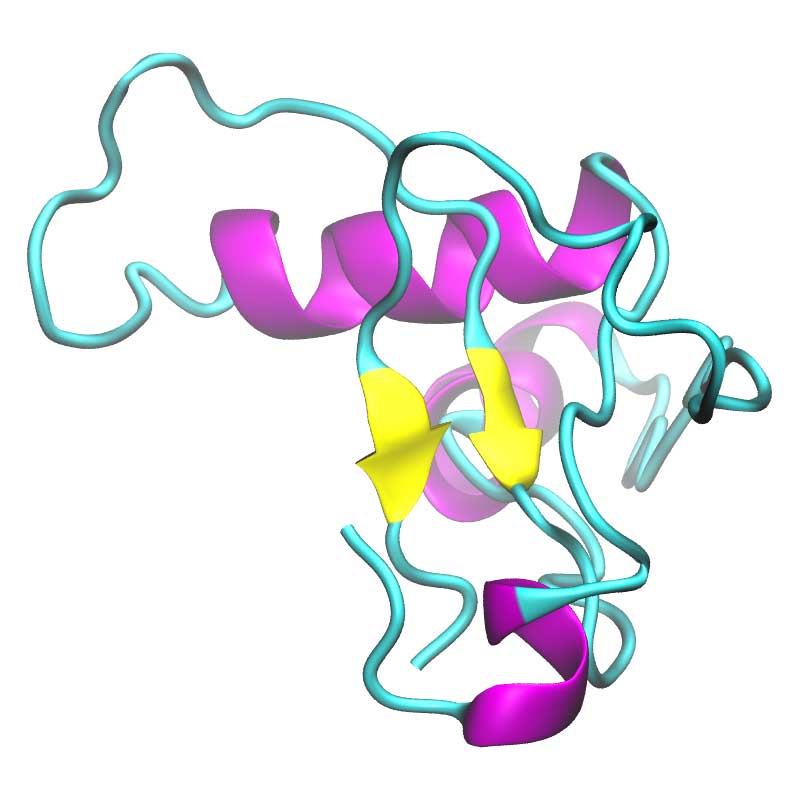
The specific role of nsp2 has not been established; it may be that nsp2 assists other viral proteins in performing their function, such as regulating the autophagy defense response or promoting mitochondrial dysfunction, thereby helping viral replication or effecting disease severity. Some evidence links nsp2 to mitochondrial dysfunction and autophagy via prohibitin 1 (PHB1), PHB2 and LC3.
The nonstructural protein 3 (nsp3) acts as a phosphatase and its catalytic domain is conserved, sharing homology with yeast, archaea, and E. coli. The papain-like protease domain of nsp3 is implicated in inhibiting components of NF-κB, interferon-beta, and p53, thereby disrupting the host immune response. A substitution in SARS-CoV-2 likely intensifies the interaction with host interferon-stimulated gene products.
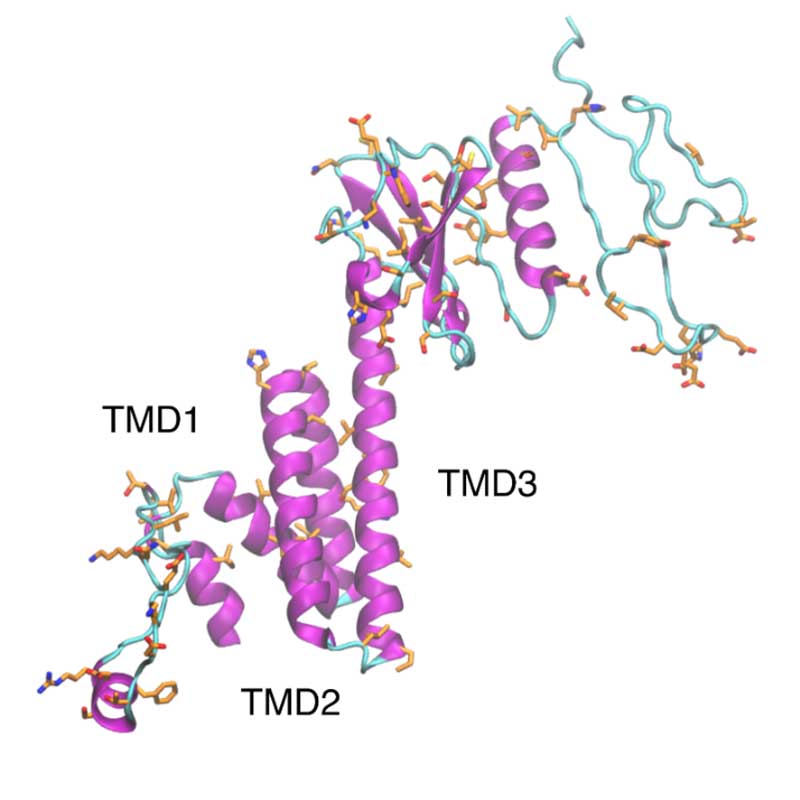
The nonstructural protein 4 (nsp4) is essential for membrane rearrangements during viral replication in a mechanism involving nsp3. Nsp4 is thought to be a tetra spanning transmembrane protein and disruption in glycosylation sites within the luminal loop give rise to aberrant double membrane vesicles. Structural information about nsp4 is scarce, thus a low-resolution model was generated for the full nsp4, using the ab initio protocol. The sequence of SARS-CoV-2 nsp4 is 80% identical to SARS-CoV-1 nsp4. However, non-conservative substitutions may affect the arrangement and packing of transmembrane segments.
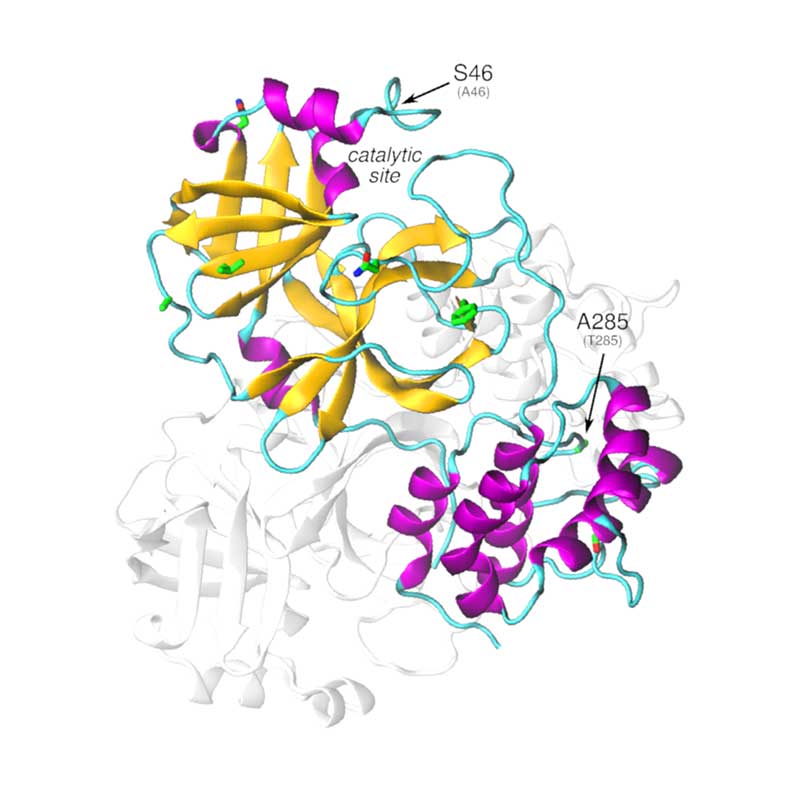
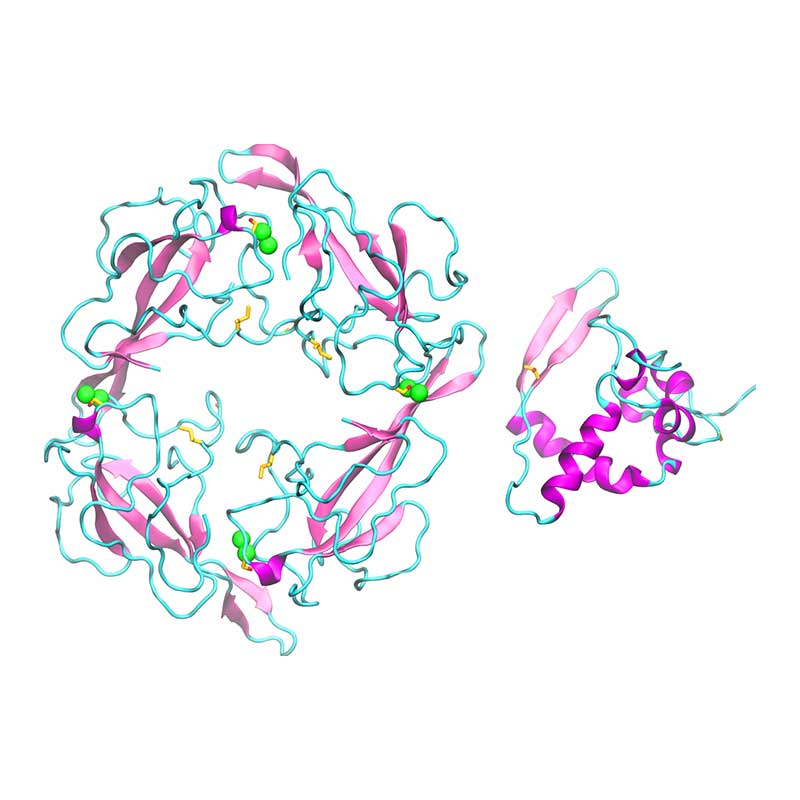
The nonstructural protein 5 (nsp5, also known as 3CLpro), is the main protease of the coronavirus genome, exhibiting as main known role the cleavage of the polyproteins translated from the viral RNA viral. Dimerization of SARS-CoV-1 3CLpro is essential to stabilize the catalytic site. The dimer interface is highly conserved within SARS-CoV-1 and -CoV-2, however, substitutions may affect dimer interaction and phosphorylation pattern of 3CLpro in SARS-CoV-2. Given that the substrate binding site of SARS-CoV-2 3CLpro is very similar to PEDV 3CLpro, it is possible that SARS-CoV-2 3CLpro is also active towards NF-kB essential modulator, thus suppressing host immune response.
Nsp6, along with nsp3 and nsp4, plays a critical role in membrane rearrangement. In SARS-CoV-1 nsp6 is known to activate autophagy by inducing perinuclear vesicles localized around the microtubule organization center. SARS-CoV-2 nsp6 is 87% identical to SARS-CoV-1 nsp6. Non-conservative mutations are located on the protein surface.
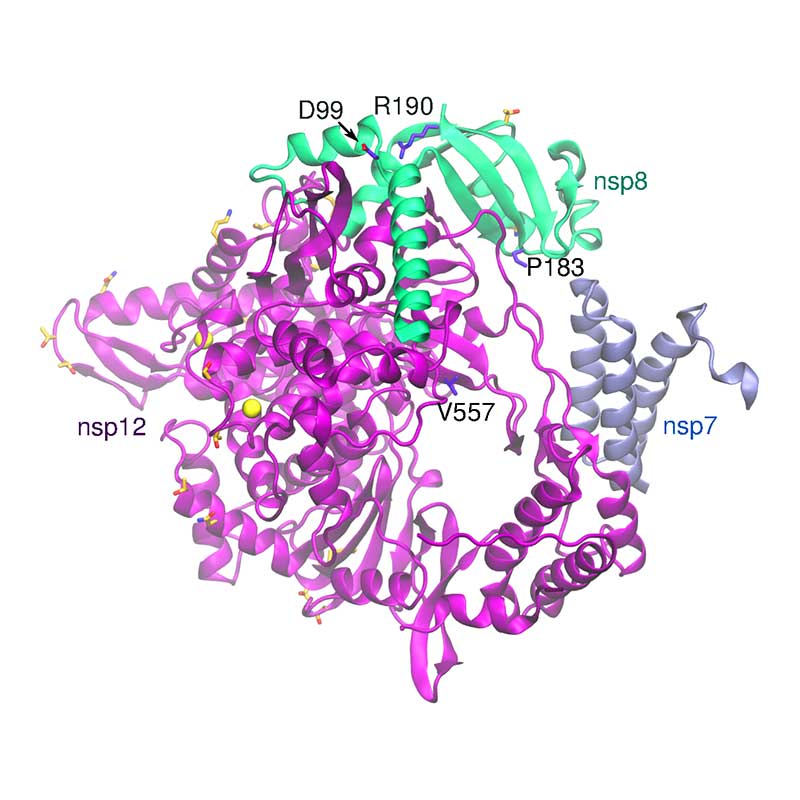
The complex formed by nsp7, 8, 9, 10, and 12 is responsible for the replication and transcription of the viral genome. Nsp12 encodes the RNA-dependent RNA polymerase (RdRp) domain. Nsp7 forms a hexadecameric complex with nsp8 that may act as a processivity clamp for the RNA polymerase. When compared to SARS-CoV-1, key functional residues are fully conserved in SARS-CoV-2, while all non-conservative substitutions are located in the complex surface. Several of these substitutions involve residues that are potential sites for post-translational modifications, indicating a variation in post-translational patterns relative to the SARS-CoV-1 RNA polymerase complex.
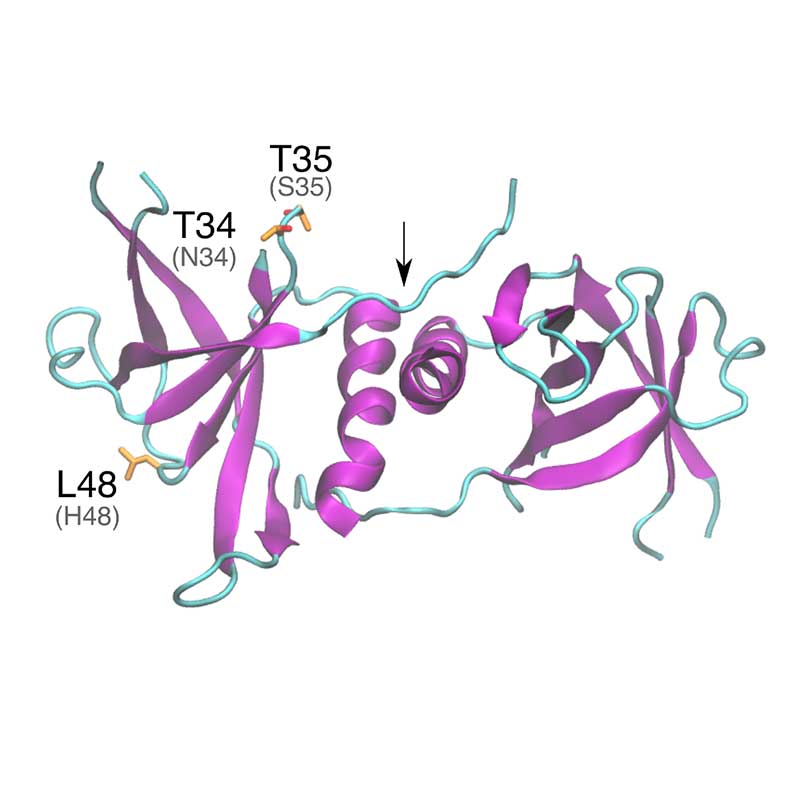
Nsp9 is able to form dimers that bind either ssDNA and ssRNA and is thought to protect the coronavirus genome from degradation during replication. Deletion of nsp9 in the mouse β-coronavirus (hepatitis virus , MHV), impairs viral RNA synthesis and viral infection. SARS-CoV-1 and SARS-CoV-2 nsp9 sequences are highly conserved, but substitutions with potential impact in post-translational modifications were verified.
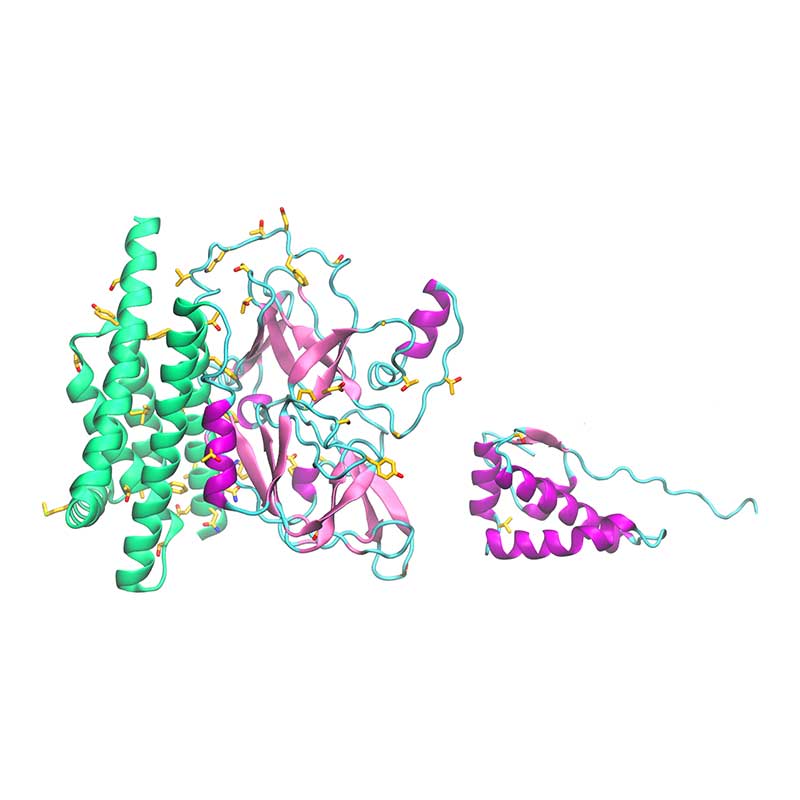
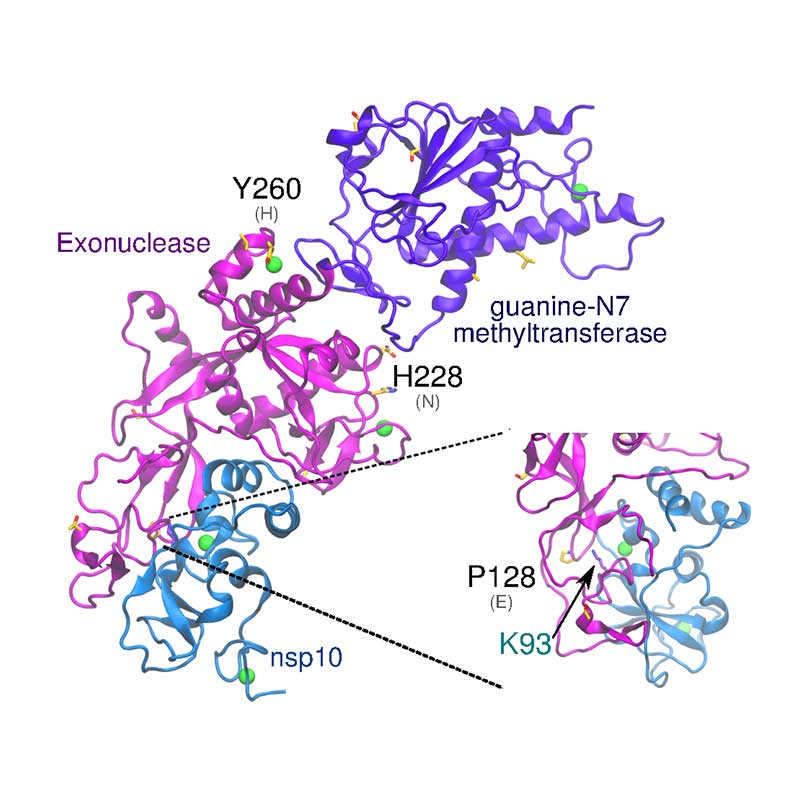
The exonuclease domain of nsp14 is imperative for replication fidelity within RNA viruses and has been shown to function as a proofreading exoribonuclease. Nsp14 associates with nsp10 that provides the ability to excise mismatched nucleotides, and disruption of this heterodimer was shown to decrease replication fidelity. SARS-CoV-1 and -CoV-21 nsp10 proteins are highly conserved (identity 97%), with no non-conservative substitutions between them. SARS-CoV-2 nsp14 is 73% identical to the SARS-CoV-1 counterpart. The substitution Glu128Pro in nsp14 is found close to the interface with nsp10, potentially affecting binding of nsp10-nsp14.
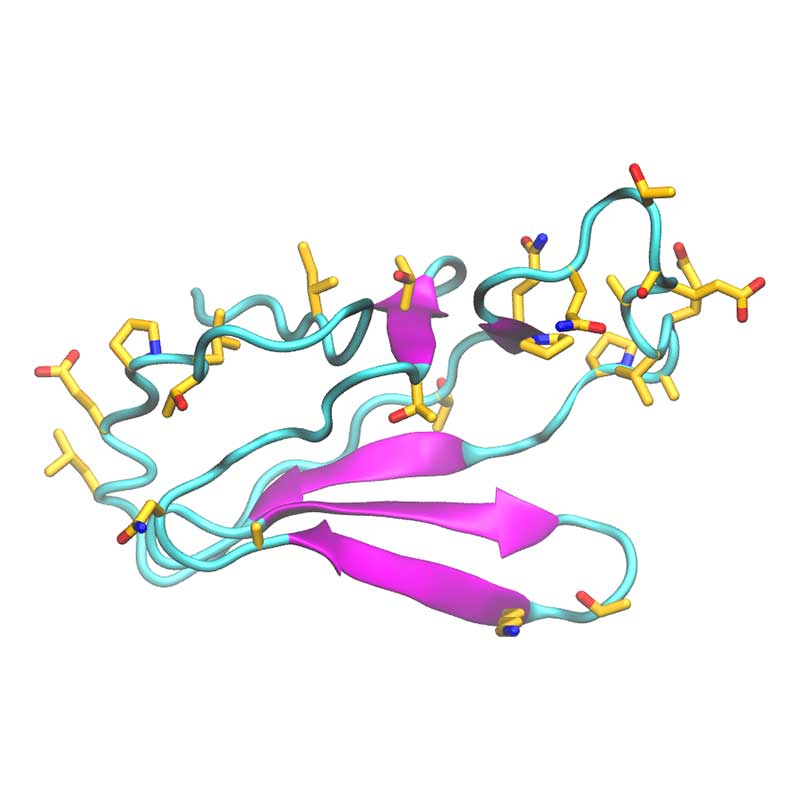
Nsp11 is a short peptide (13 a.a.) and is cleaved from both polyproteins pp1a and pp1ab. In SARS-CoV-1 nsp11 has been implicated in RNA synthesis. SARS-CoV-2 nsp11 has 85% sequence identity to SARS-CoV-1 nsp11.
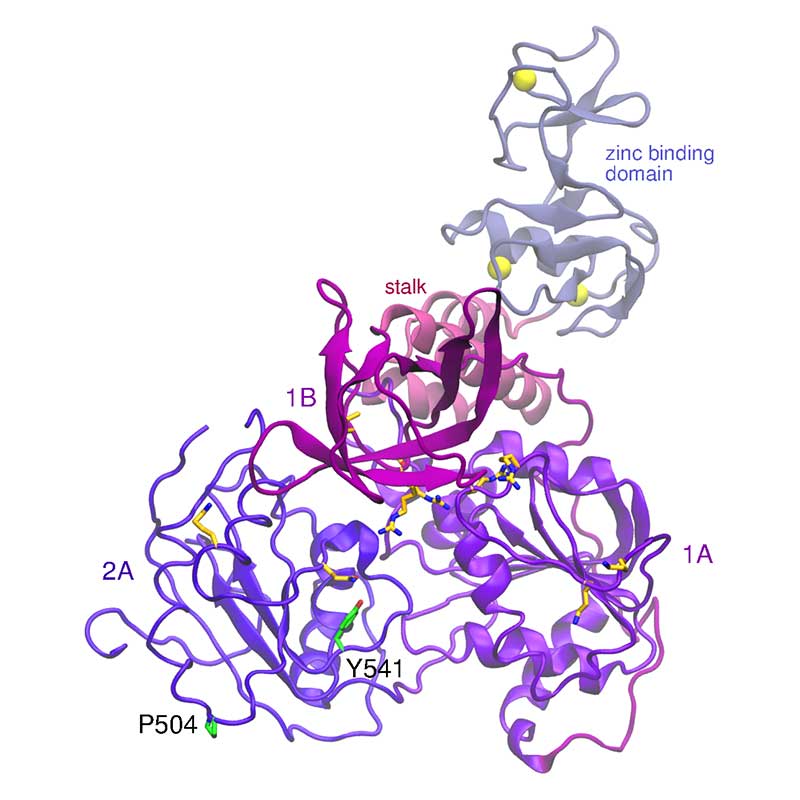
The nonstructural protein 13 (nsp13) is involved in a number of functions, such as NTPase, dNTPase, RTpase, RNA helicase, and DNA helicase activity. A single mutation in SARS-CoV-1 nsp13 stalk domain, resulted in a dramatic decrease in viral infectivity in vitro. The essential nature of nsp13 in the viral replication cycle, and its multifunctional nature has made it an attractive target for vaccine research. SARS-CoV-2 nsp13 is fully conserved relative to SARS-CoV-1.
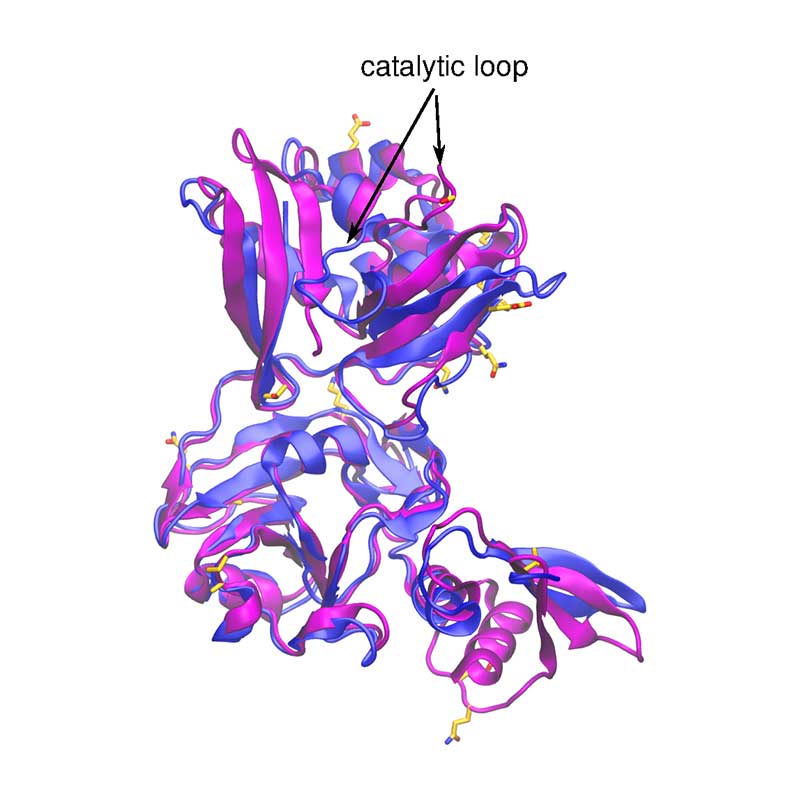
Nsp15 is a Nidoviral RNA uridylate-specific endoribonuclease and its C-terminal is a catalytic domain belonging to the EndoU family of enzymes. Within SARS-CoV-1 and -CoV-2, nsp15 is very conserved (89 % identity).
The nonstructural protein 16 (nsp16) is involved in capping of viral mRNA to protect it from host degradation, and it has been demonstrated that it has to be associated with nsp10 to be active. Mutation that increases hydrophobic interaction between nsp10 and the hydrophobic pocket of nsp16, increases methyltransferase activity of nsp16. Conversely, a mutation in the RNA binding site of nsp16, completely abolished its methyltransferase activity. All these key amino acids are conserved in SARS-CoV-2 nsp16, but non-conservative substitutions in their vicinity may have a steric effect.
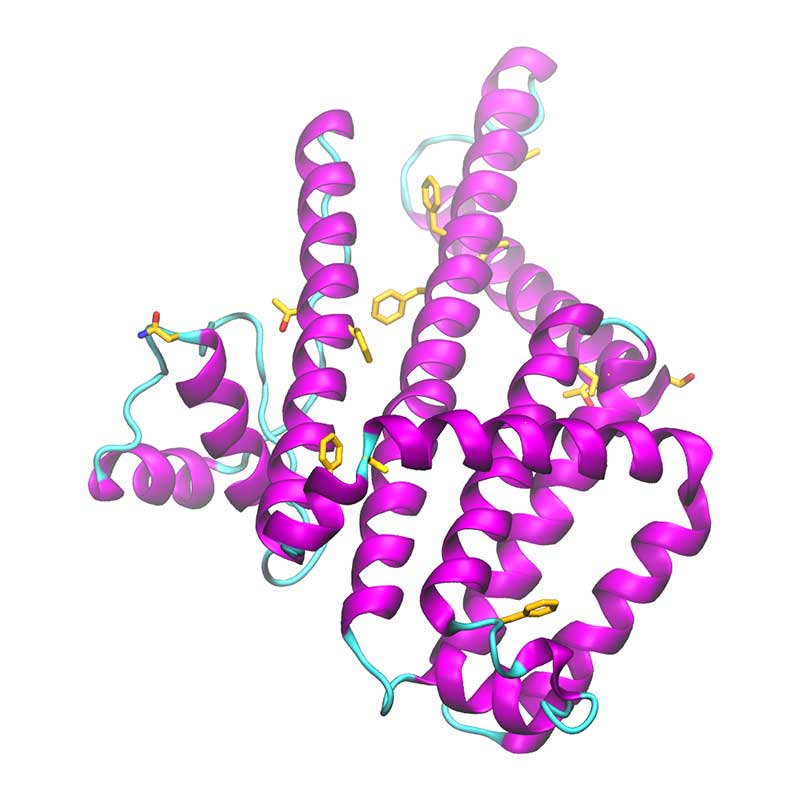
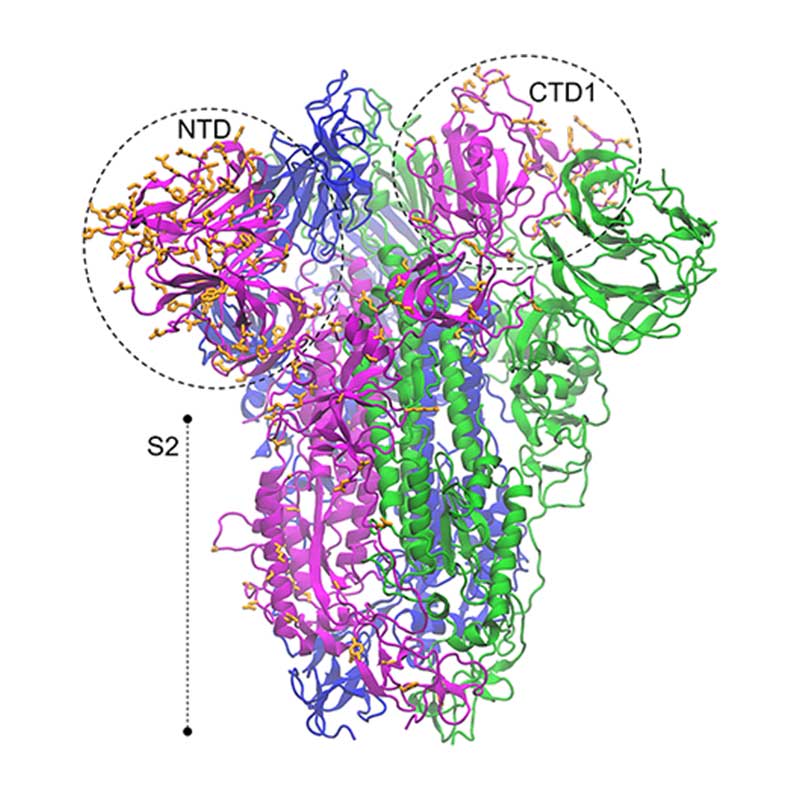
The spike glycoprotein (S) is the main promoter of virus entry into host cells. These highly glycosylated proteins protrude from the viral surface to interact with the host cell receptor(s). The receptor binding domain of SARS-CoV-2 S harbors many non-conservative substitutions relative to SARS-CoV-1 S. Understanding structure-dynamics relationships in S-receptor complexes can be of key importance to develop effective drugs targeting this interaction and to unveil the pathways of infection by SARS-CoV-2.
ORF3a is an integral transmembrane protein, localized mostly in the Golgi complex, cytoplasm and cell surface. ORF3a is able to form homotetramers with ion channel properties. Regarding pathogenic effects, ORF3a is linked to inflammatory responses, weak INF responses, innate immunity responses, trigger apoptosis and modulate cell cycle. Non-conservative mutations with a potential impact in apoptosis, cell cycle arrest, caveoline-1 binding, phosphorylation, glycosylation and RNA binding were verified.
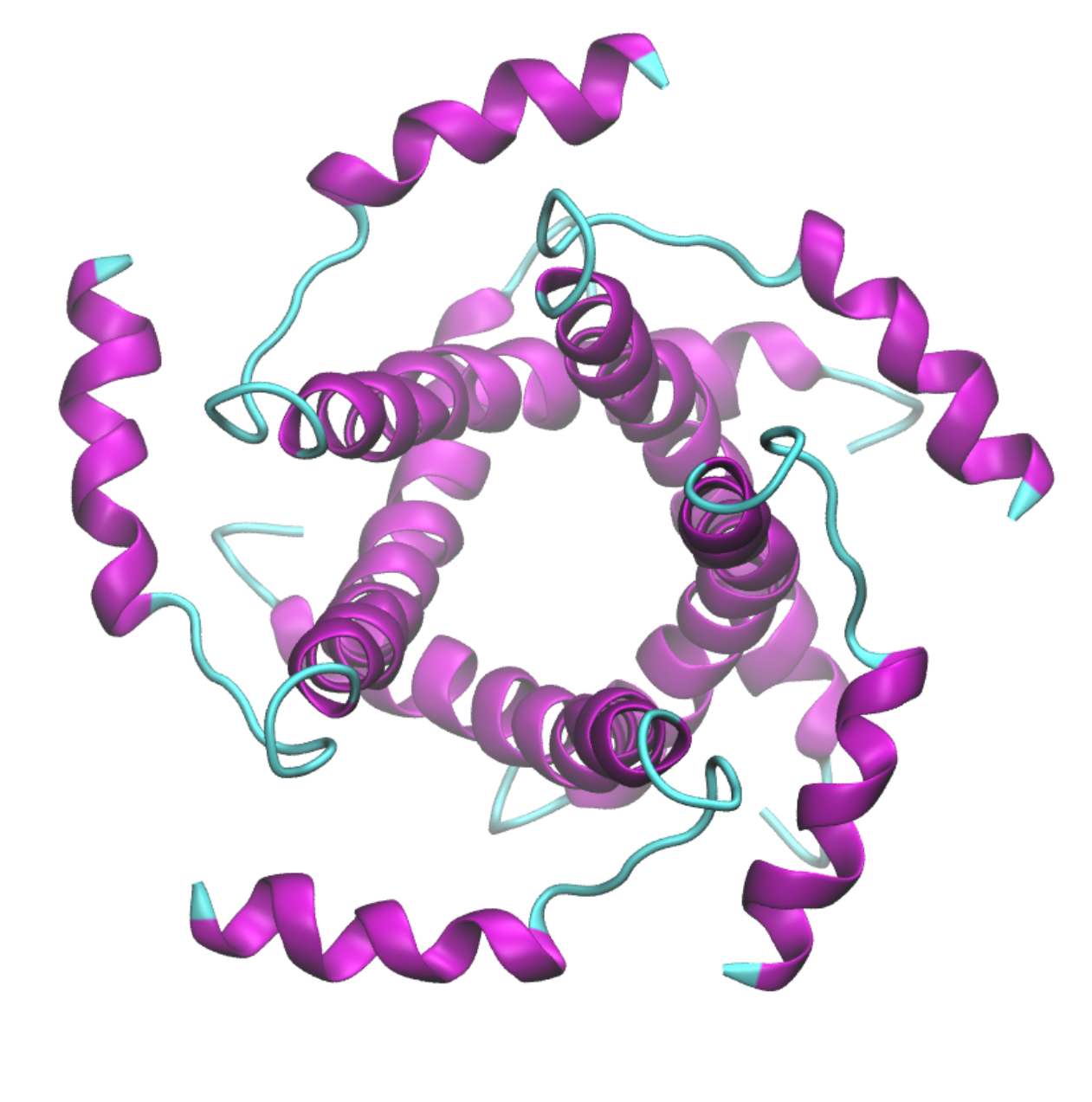
E protein is a type 1 transmembrane protein able to form pentamers creating pores with ion transport activity. E protein localizes mainly in the ER and Golgi apparatus, where it participates in assembly, budding and intracellular trafficking of newly formed virions. E protein is involved in tight junctions disruption in the lungs to reach the alveolar wall and develop into a systemic infection, triggering an overexpression of inflammatory cytokines and lymphopenia. Four variations in SARS-CoV-2 E protein relative to SARS-CoV-1 were verified, all located in the C-terminal to the cytoplasmic face, thereby potentially modulating E protein interaction with other proteins.
Membrane (M) protein is the most abundant of all coronavirus structural proteins. M protein main functions are membrane curvature initiation, RNA packing and viral particle budding. Besides, M protein has been linked to apoptosis, suppression of inflammatory responses and IFN signaling. Most of the variation between SARS-CoV and SARS-CoV-2 M occurs in the cytoplasmic domain at the C-terminus, involving several sites that can potentially undergo post-translational modification. Therefore, these mutations could affect M protein function and antigenic properties.
ORF6 localizes in the perinuclear and endoplasmic reticulum. ORF6 is known to interact with the viral proteins ORF9b and nsp8, suggesting ORF6 may play a role in virus replication. Besides, ORF6 has been associated with impairing the activation of IFN signaling and membrane rearrangements. SARS-CoV-1 and SARS-CoV-2 ORF6 proteins are 68.85% identical, with most of the substitutions located in the C-terminus helix. Among them, the introduction of new putative ubiquitination sites and mutations in a critical area for ORF6 function were verified.
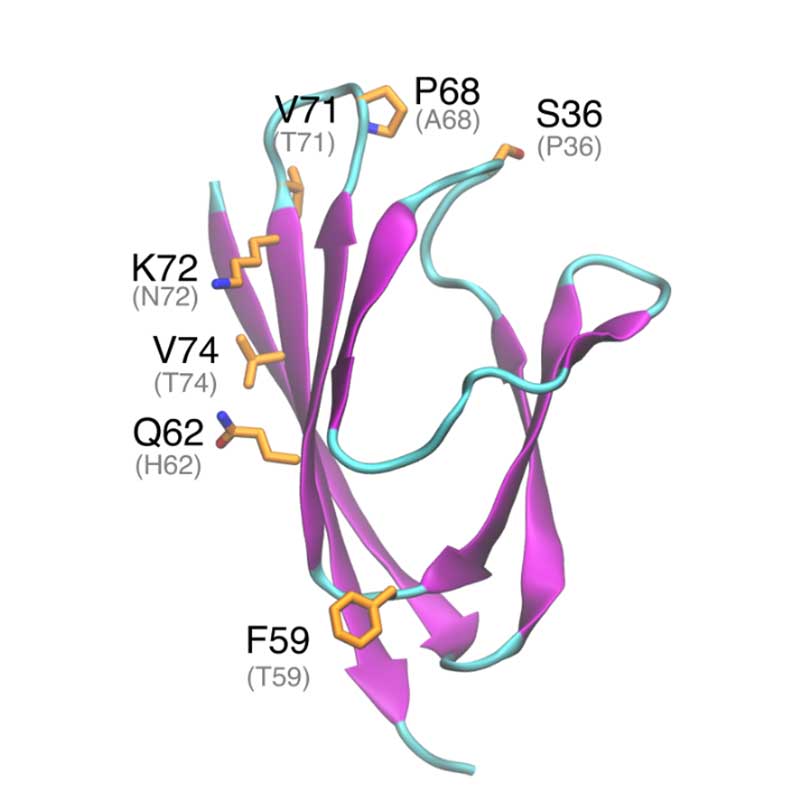
ORF7a is a type I transmembrane protein, localized mainly in ER, Golgi apparatus and in the cell surface. ORF7a is able to prevent virus tethering at the plasma membrane by BTS-2. Further in vitro experiments showed that ORF7a is able to induce apoptosis in a caspase dependent manner. ORF7a protein is very conserved between SARS-CoV-1 and SARS-CoV-2 with 85% sequence identity. The substitutions may affect ORF7a interaction with other viral or host proteins.
This accessory protein is an integral transmembrane protein and it localizes in the cis- and trans- Golgi. MSARS-CoV-2 ORF7b is 81 % identical to SARS-CoV-1 ORF7b. All substitutions are found in the terminals, being the transmembrane helix fully conserved.
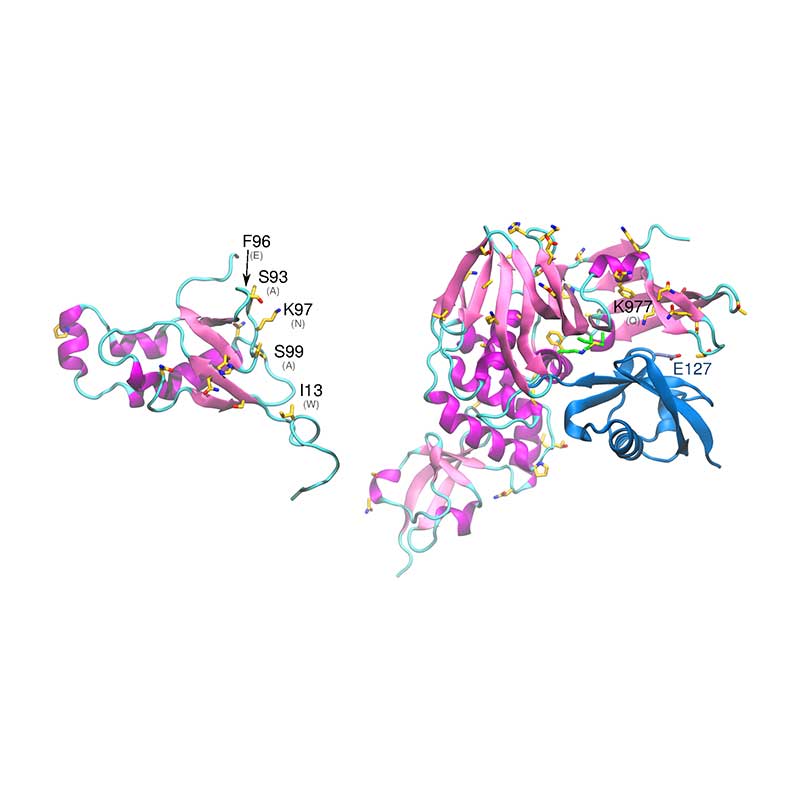
ORF8 is one of the most variable among SARS-CoV-1. Remarkably, 29 nucleotide N-terminal sequence has gone under gradual deletion during SARS-CoV-1 spread in humans. Early tests showed a decrease in viral replication by up to 23-fold when this sequence was removed. SARS-CoV-2 ORF8 is 127 amino acids long and 45% identical to SARS-CoV-1 ORF8b.
Nucleocapsid (N) protein packages the viral RNA into a helical ribonucleocapsid and plays a key role in viral assembly. SARS-CoV-2 N protein is a 419 amino acid long protein and is 89% identical to SARS-CoV-1 nucleocapsid. The RNA binding domain of N is located at the N-terminal and all these residues are fully conserved in SARS-CoV-2 N.
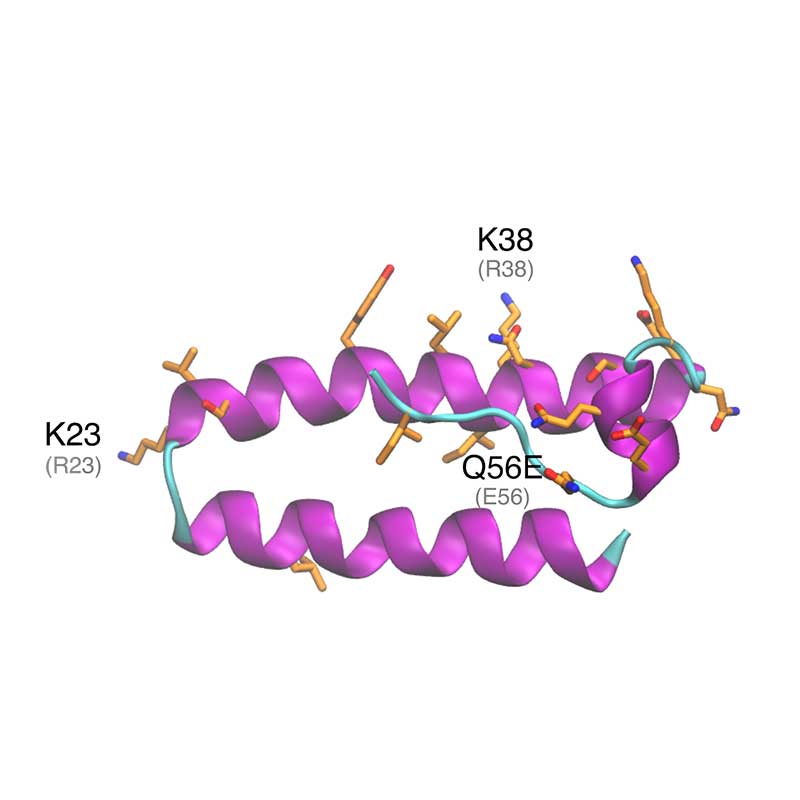
ORF9b is an accessory protein synthesized from an alternative reading frame in the N gene. At mitochondrial level, ORF9 promotes suppression of the activation of INF regulatory factors and NF-κB. ORF9 forms a symmetric dimer where the monomer interactions resemble a handshake. Dimer assembling creates a hydrophobic tunnel that can accomodate a long fatty acid chain. Thus, ORF9 could anchor itself to a lipidic membrane by internalizing one or more lipidic tails. SARS-CoV-2 ORF9 shares high homology with SARS-CoV-1 (72·45% protein identity).
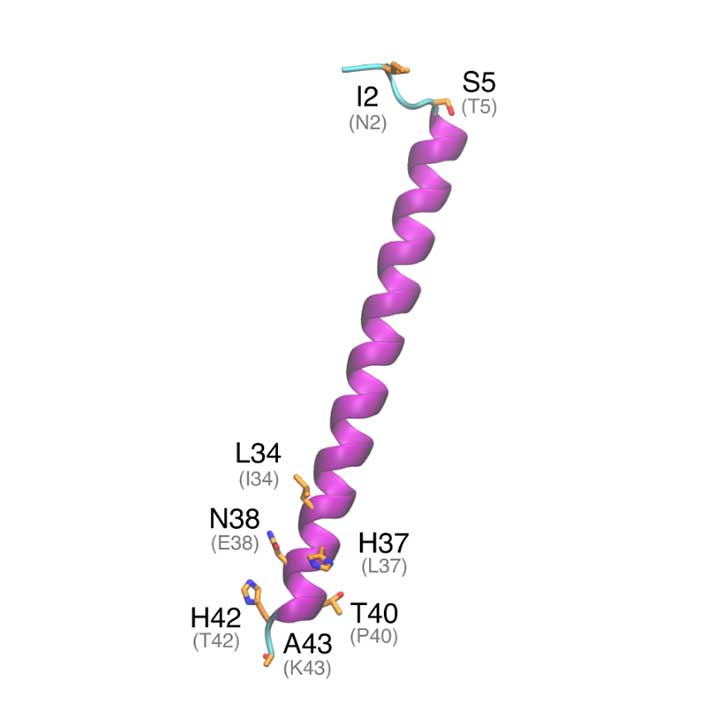
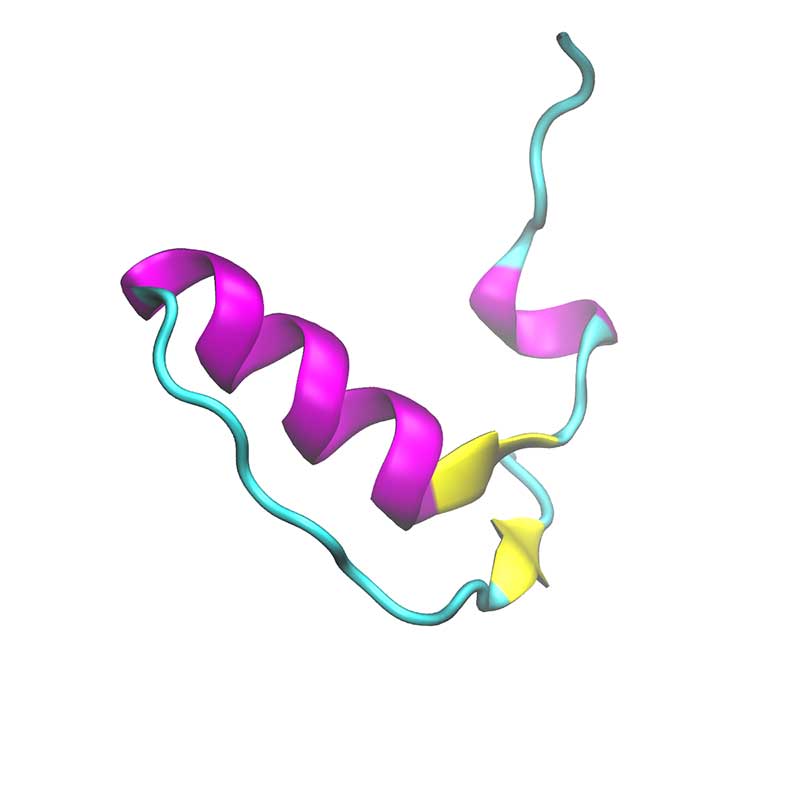
Within SARS-COV-2, ORF10 is thought to not have any functional protein purpose as it was not found within sub genomic mRNA sequencing. In another study, ORF10 might hijack components of the ubiquitination machinery and degradation. ORF10 of SARS-COV-2 is the last and the shortest predicted coding sequence upstream of the poly-A tail. ORF10 is predicted to harbor a long helix and then parallel ϐ-sheets. ORF10 is not found within the SARS-CoV-1 proteome, making it unique to SARS-COV-2.